We (Julie) are still very active in the long-read sequencing business. Please see an updated figure regarding the output per P2 flow cell below. Reach out if you would like to collaborate!
#PlantGenomics
🌱Takeaway: Ecological isolation, limited dispersal, and life-history traits drive both speciation and extinction risk in Homoranthus. (9/9)
#PlantScience #EvolutionaryBiology #PopulationGenetics #PlantGenomics #AoBpapers
🌿🌍 Join us to learn about the newly published “Geographically proximate rare species exhibit strong population divergence while maintaining intraspecific genetic diversity in Homoranthus (Myrtaceae)” in @AnnBot by Eilish McMaster and co-authors. (1/9)
#PlantScience #EvolutionaryBiology #PopulationGenetics #PlantGenomics #AoBpapers
Celebrate the brilliance of women driving plant sciences forward at Pucker Lab on International Day of Women and Girls in Science! 🌱🔬 Here's to curiosity, innovation, and shattering glass ceilings in STEM! 💪♀️ #WomenInScience #PuckerLab #PlantGenomics #GirlsInSTEM
Just 4 days of Python and our students are extracting insights from plant genomics data — all without AI! 🌱
Learn Python & bioinformatics in Bonn:
https://www.izmb.uni-bonn.de/en/pbb/contact
#Python #Bioinformatics #PlantGenomics #ScienceEducation #Genomics
@PuckerLab
@samnm
New year, new students! ✨
We are pleased to welcome Maria, who is going to be with us for a short time to learn everything you need to start a plant genomics project, from DNA extraction to sequencing, and of course we use ONT for our projects, with Julie teaching all new students about the technology! 🧬
Our Functional Annotation course has come to an end, and we are happy with how it went. More than 20 early-career researchers explored plant genomics together and brought amazing curiosity to every session. Grateful to all participants—your passion and hard work give us real optimism for the next generation of plant science. 🌱🧬
#PlantGenomics #Bioinformatics #FunctionalAnnotation #ScienceEducation
Our course “From Gene Models to Biological Insights – Functional Annotation” has officially concluded! Proud to have supported 20+ early-career researchers in developing their skills in plant genomics. 🌱🧬
Looking forward to seeing their discoveries!
#PlantGenomics #Bioinformatics #FunctionalAnnotation #ScienceEducation
Our paper “Giant chromosomes of a tiny plant” is out in
@GigaScience
! We present a T2T genome assembly of the simple thalloid liverwort Apopellia endiviifolia with truly giant chromosomes. 🌱🧬 #plantgenomics #T2T #liverworts #nanoporeSequencing
@nanopore
tinyurl.com/m8tkkxd3
Preparing for our #PlantGenomics courses! 🌿🧬
Spots are nearly filled—check our website if you would like to participate and learn about long-read sequencing + cloud-based data analysis.
Boosts appreciated!
🔗 https://www.izmb.uni-bonn.de/en/pbb/news#denbi2025
🌿 Interested in #PlantGenomics?
Learn the basics of genome sequence assembly and annotation in our upcoming courses in Bonn!
🔗 https://www.izmb.uni-bonn.de/en/pbb/news#denbi2025
#Genomics #Bioinformatics #DeNBI #ScienceEducation
@PuckerLab @deNBI
The Plant Genomics & Bioinformatics seminar is starting this winter at the University of Bonn 🌿
Dive into genome analysis, computational tools, and modern plant research:
🔗 https://github.com/bpucker/teaching
#PlantGenomics #Bioinformatics #OpenScience #UniBonn #DataLiteracy
@PuckerLab
Excited to share our new #preprint!
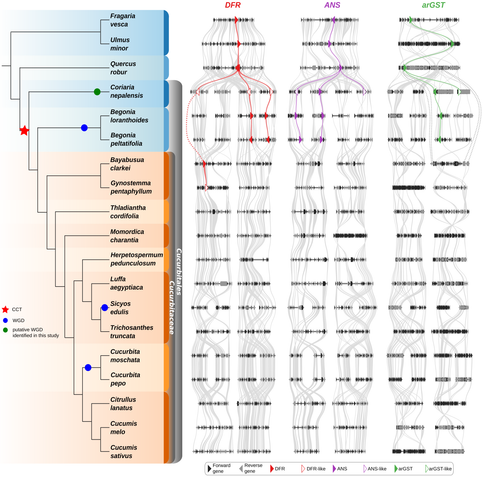
Large-scale Phylogenomics Reveals Systematic Loss of Anthocyanin Biosynthesis Genes at the Family Level in #Cucurbitaceae
🔗 https://doi.org/10.1101/2025.10.06.680802
We uncover family-level losses of anthocyanin and proanthocyanidin biosynthesis genes — shedding light on the evolution of pigment diversity in cucurbits.
#Phylogenomics #Evolution #PlantGenomics #Anthocyanins #OpenScience
@PuckerLab
🌹This work highlights how hybridization and a single key mutation shaped roses' beauty and diversity. (9/9)
#Botany #AoBpapers #PlantScience #PlantGenomics #RoseGenetics
🎉 Good news! The paper ‘Molecular investigation of the progenitors, origin, and domestication patterns of diploid Chinese old garden roses’ in @AnnBot by Cheng Zhang and co-authors is now #free for a limited time 🧵(1/9)
👉 https://doi.org/p8g9
#Botany #AoBpapers #PlantScience #PlantGenomics #RoseGenetics
🧬 New PCI Genomics recommendation! HaploCharmer: a Snakemake workflow for read-scale haplotype calling adapted to polyploids, by Simon Rio and colleagues.
✨ Recommended by Lucia Campos Dominguez who praised its scalability for highly complex polyploids
📄 Preprint: https://doi.org/10.1101/2025.03.14.642807
🔗 Recommendation: https://doi.org/10.24072/pci.genomics.100434
🌴This discovery sheds light on how sex chromosomes evolve in plants and shows that evolution can sometimes take the same path twice. (9/9)
🌱 CRISPR breakthrough! Editing GSL5 gene provides broad-spectrum clubroot resistance in cruciferous crops. Genetic game-changer for agricultural sustainability! 🚜 #PlantGenomics #CRISPRrevolution #CropProtection https://emmecola.github.io/genomics-daily
Computational Postdoctoral Fellow
Joint Genome Institute, Lawrence Berkeley National Laboratory
See the full job description on jobRxiv: https://jobrxiv.org/job/joint-genome-institute-lawrence-berkeley-national-laboratory-27778-computational-postdoctoral-fellow/
#aiforscience #largelanguagemodels #plantgenomics #ScienceJobs #hiring #research
https://jobrxiv.org/job/joint-genome-institute-lawrence-berkeley-national-laboratory-27778-computational-postdoctoral-fellow/?fsp_sid=1796
Postdoc opportunity with @tomasbruna.bsky.social here at @jgi.doe.gov @berkeleylab.lbl.gov: Develop, benchmark, and apply Plant Genomic Language Models (gLMs) 🌿🧬💻 jobs.lbl.gov/jobs/computa... #AI #PlantGenomics #gLMs #Postdoc